In HEP analyses, we constantly face the challenge of extracting signal yields, measuring particle properties, and determining background shapes from experimental data. Whether we're measuring a branching fraction, searching for a rare decay, or extracting a resonance mass, you need to fit probability distribution functions (PDFs) to our data.
But how do we do this? The two most common approaches are binned and unbinned maximum likelihood fits. The choice between binned and unbinned maximum likelihood fits isn't merely academic, it directly impacts our statistical sensitivity, systematic uncertainties, and how much resources (i.e. CPU cycles) we need to allocate/waste for the analysis.
At the LHC, where we routinely analyze datasets with millions of events (viva Amplitude analyses), this choice matters. A naive binned fit might throw away information through poor binning choices, while an unbinned fit on truly massive datasets could be computationally prohibitive. So let me walk through the theoretical foundations and practical implementations of both methods, using realistic examples from invariant mass analyses.
Mathematical Framework
The Likelihood Function
Given observations drawn from a probability density function parametrized by , our goal is to estimate by maximizing the likelihood.
Unbinned Likelihood:
The unbinned likelihood is straightforward, it's simply the product of PDF values evaluated at each observed data point:
In practice, we work with the negative log-likelihood (NLL) for numerical stability:
Binned Likelihood:
For binned data, we partition the observable space into bins. Let be the observed count in bin , and be the expected count:
The binned likelihood follows Poisson statistics in each bin:
The corresponding NLL becomes:
Since doesn't depend on , we typically drop it during minimization.
Statistical Efficiency
The asymptotic efficiency of binned vs unbinned fits depends on the binning granularity. As the number of bins with bin width , the binned likelihood converges to the unbinned one. However, for finite bins, we lose information.
The key insight here is that unbinned fits are statistically optimal. They extract all available information from the data. Binned fits introduce an effective loss of information, which can be quantified through the loss in Fisher information. For a well-chosen binning scheme with sufficient statistics per bin, the efficiency loss is typically small (few percent). However, poor binning can significantly degrade your sensitivity.
Invariant Mass Analysis
Let's implement both methods for a semi-realistic common HEP scenario, measuring a resonance (signal) over a combinatorial background in an invariant mass spectrum. This is actually the most common thing we do in HEP, and it's a good example to illustrate the differences between binned and unbinned fits.
The first thing we would do is to set up our environment and dependencies. We will use standard numerical libraries and will not resort to ROOT. We would use Numpy for numerical operations, Matplotlib for plotting, and SciPy because we don't want to re-write all statistical functions from scratch. I also define good plotting styles (because why not?). It is also important to set a random seed for reproducibility, especially when generating MC samples.
The code to do this is as follows:
import numpy as np
import matplotlib.pyplot as plt
from scipy import stats
from scipy.optimize import minimize
from scipy.integrate import quad
import warnings
warnings.filterwarnings('ignore')
# Set style for good-quality plots
plt.rcParams['figure.figsize'] = (12, 8)
plt.rcParams['font.size'] = 12
plt.rcParams['axes.labelsize'] = 14
plt.rcParams['axes.titlesize'] = 14
plt.rcParams['xtick.labelsize'] = 12
plt.rcParams['ytick.labelsize'] = 12
plt.rcParams['legend.fontsize'] = 11
# Set random seed for reproducibility
np.random.seed(42)
Define PDF Components
Now let's define the PDFs for our signal and background. We'll use a Crystal Ball function for the signal, which is a common choice in HEP for modeling resonances with radiative tails. For the background, we'll use an exponential decay, which is typical for combinatorial backgrounds. Although it is not going to be good if you are trying simultaneous fits with multiple components, it is a good starting point for many analyses.
For our signal, we'll use a Crystal Ball function for modeling resonances with radiative tails:
For the background, we'll use an exponential decay, typical of combinatorial backgrounds:
Now let's implement these two functions and the total PDF that combines both signal and background components. The total PDF will be a weighted sum of the signal and background PDFs, normalized to integrate to 1 over the specified range.
def crystal_ball(x, mu, sigma, alpha, n):
"""
Crystal Ball function
Parameters:
-----------
x : array-like or float
Observable (invariant mass)
mu : float
Peak position
sigma : float
Core resolution
alpha : float
Tail parameter (typically ~1-2)
n : float
Tail exponent (typically ~2-5)
"""
# Ensure x is always treated as an array internally
x_array = np.atleast_1d(x)
t = (x_array - mu) / sigma
# Gaussian core
abs_alpha = np.abs(alpha)
# This calculates A and B coefficients
A = (n / abs_alpha) ** n * np.exp(-0.5 * alpha**2)
B = n / abs_alpha - abs_alpha
# Now, compute the function
result = np.zeros_like(x_array, dtype=float)
# Here is the Gaussian part
gaussian_region = t > -abs_alpha
result[gaussian_region] = np.exp(-0.5 * t[gaussian_region]**2)
# And a Power-law tail
tail_region = t <= -abs_alpha
result[tail_region] = A * (B - t[tail_region])**(-n)
# If the original input was a scalar, return a scalar value
if np.isscalar(x):
return result.item()
return result
def exponential_bkg(x, lamb, x_min):
"""
Exponential background - for a combinatorial backgrounds
Parameters:
-----------
x : array-like
Observable
lamb : float
Decay constant
x_min : float
Minimum of range (normalization point)
"""
return lamb * np.exp(-lamb * (x - x_min))
def total_pdf(x, n_sig, n_bkg, mu, sigma, alpha, n, lamb, x_min, x_max):
"""
Total PDF: signal + background
Returns normalized PDF (integrates to 1)
"""
# Crystal Ball signal (which needs to normalize)
sig = crystal_ball(x, mu, sigma, alpha, n)
# Numerical normalization for Crystal Ball
norm_sig, _ = quad(lambda t: crystal_ball(t, mu, sigma, alpha, n),
x_min, x_max, limit=100)
sig = sig / norm_sig
# Exponential background (normalized)
bkg = exponential_bkg(x, lamb, x_min)
norm_bkg = 1 - np.exp(-lamb * (x_max - x_min))
bkg = bkg / norm_bkg
# Total event count
n_total = n_sig + n_bkg
# Weighted sum
return (n_sig * sig + n_bkg * bkg) / n_total
But we also want to normalize the PDFs so that they integrate to 1 over the specified range. This is important for both the signal and background components, as it allows us to interpret the PDFs as probability distributions. We can achieve this by integrating the PDFs over the range of interest and dividing by the integral.
def signal_pdf_normalized(x, mu, sigma, alpha, n, x_min, x_max):
"""Normalized signal PDF"""
sig = crystal_ball(x, mu, sigma, alpha, n)
norm, _ = quad(lambda t: crystal_ball(t, mu, sigma, alpha, n),
x_min, x_max, limit=100)
return sig / norm
def background_pdf_normalized(x, lamb, x_min, x_max):
"""Normalized background PDF"""
bkg = exponential_bkg(x, lamb, x_min)
norm = 1 - np.exp(-lamb * (x_max - x_min))
return bkg / norm
MC Sample Generation
Now that we have our PDFs defined, we can generate some MC samples for both signal and background. We'll use accept-reject sampling for the Crystal Ball signal and inverse transform sampling for the exponential background. The reason we use accept-reject for the Crystal Ball is that it has a non-trivial shape, while the exponential is straightforward to invert.
Here is the python code to generate these samples:
def generate_signal_events(n_events, mu, sigma, alpha, n, x_min, x_max, max_attempts=10):
"""
Generate signal events using accept-reject sampling
This mimics how we'd generate MC for detector simulation studies
"""
events = []
attempts = 0
while len(events) < n_events and attempts < max_attempts:
# Proposal distribution: uniform over range
n_propose = int(n_events * 2) # Oversample
x_propose = np.random.uniform(x_min, x_max, n_propose)
# Evaluate PDF
pdf_vals = crystal_ball(x_propose, mu, sigma, alpha, n)
pdf_max = np.max(pdf_vals) * 1.1 # Adding 10% margin for safety
# Accept-reject
u = np.random.uniform(0, pdf_max, n_propose)
accepted = x_propose[u < pdf_vals]
events.extend(accepted)
attempts += 1
return np.array(events[:n_events])
def generate_background_events(n_events, lamb, x_min, x_max):
"""
Generate exponential background events using inverse transform sampling
"""
u = np.random.uniform(0, 1, n_events)
# Inverse CDF of exponential
x = x_min - np.log(1 - u * (1 - np.exp(-lamb * (x_max - x_min)))) / lamb
return x
# Define our "true" parameters (what we'll try to extract)
TRUE_PARAMS = {
'mu': 5.28, # B meson mass in GeV (example)
'sigma': 0.025, # ~25 MeV resolution
'alpha': 1.5, # Tail parameter
'n': 2.5, # Tail exponent
'lamb': 2.0, # Background slope
'n_sig': 1000, # Signal events
'n_bkg': 5000, # Background events
'x_min': 5.0, # Mass window min
'x_max': 5.6 # Mass window max
}
# Generate our "data"
print("Generating Monte Carlo samples...")
sig_events = generate_signal_events(
TRUE_PARAMS['n_sig'],
TRUE_PARAMS['mu'],
TRUE_PARAMS['sigma'],
TRUE_PARAMS['alpha'],
TRUE_PARAMS['n'],
TRUE_PARAMS['x_min'],
TRUE_PARAMS['x_max']
)
bkg_events = generate_background_events(
TRUE_PARAMS['n_bkg'],
TRUE_PARAMS['lamb'],
TRUE_PARAMS['x_min'],
TRUE_PARAMS['x_max']
)
# Combine into "data"
data = np.concatenate([sig_events, bkg_events])
np.random.shuffle(data) # Randomize order
print(f"Generated {len(data)} total events")
print(f" Signal: {len(sig_events)}")
print(f" Background: {len(bkg_events)}")
print(f" S/B ratio: {len(sig_events)/len(bkg_events):.3f}")
print(f" S/√B ratio: {len(sig_events)/np.sqrt(len(bkg_events)):.2f}")
We define the real parameters for our signal and background, generate the respective events, and combine them into a single dataset. We also print out some useful statistics about the generated sample. The parameters chosen here are typical for a B meson invariant mass, although it is statistically horrible for a real analysis.
When we run this code, we should see output similar to:
Generating Monte Carlo samples...
Generated 6000 total events
Signal: 1000
Background: 5000
S/B ratio: 0.200
S/√B ratio: 14.14
Visualize the Data
Now once we have generated our data, it's always a good idea to visualize it. This helps us understand the distributions and verify that our generation process worked as expected. We'll create histograms of the full dataset, as well as separate plots for the signal and background components.
# Create visualization of our data
fig, axes = plt.subplots(2, 2, figsize=(14, 10))
# Plot 1: Full data with components
ax = axes[0, 0]
bins = 60
counts, bin_edges, _ = ax.hist(data, bins=bins, alpha=0.9, color='gray',
label='Data', density=False)
ax.hist(sig_events, bins=bins, alpha=0.5, color='red',
label='Signal (truth)', density=False, histtype='step', linewidth=2)
ax.hist(bkg_events, bins=bins, alpha=0.5, color='blue',
label='Background (truth)', density=False, histtype='step', linewidth=2)
ax.set_xlabel('Invariant Mass [GeV/c²]')
ax.set_ylabel('Events / {:.1f} MeV'.format((TRUE_PARAMS['x_max']-TRUE_PARAMS['x_min'])*1000/bins))
ax.set_title('Generated Data Sample')
ax.legend()
ax.grid(alpha=0.3)
# Plot 2: Signal component zoom
ax = axes[0, 1]
mass_range = np.linspace(TRUE_PARAMS['x_min'], TRUE_PARAMS['x_max'], 1000)
sig_pdf = signal_pdf_normalized(mass_range, TRUE_PARAMS['mu'], TRUE_PARAMS['sigma'],
TRUE_PARAMS['alpha'], TRUE_PARAMS['n'],
TRUE_PARAMS['x_min'], TRUE_PARAMS['x_max'])
ax.hist(sig_events, bins=50, alpha=0.6, color='red', density=True, label='MC Signal')
ax.plot(mass_range, sig_pdf, 'r-', linewidth=2, label='True PDF')
ax.set_xlabel('Invariant Mass [GeV/c²]')
ax.set_ylabel('Normalized PDF')
ax.set_title('Signal: Crystal Ball Shape')
ax.legend()
ax.grid(alpha=0.3)
# Plot 3: Background component
ax = axes[1, 0]
bkg_pdf = background_pdf_normalized(mass_range, TRUE_PARAMS['lamb'],
TRUE_PARAMS['x_min'], TRUE_PARAMS['x_max'])
ax.hist(bkg_events, bins=50, alpha=0.6, color='blue', density=True, label='MC Background')
ax.plot(mass_range, bkg_pdf, 'b-', linewidth=2, label='True PDF')
ax.set_xlabel('Invariant Mass [GeV/c²]')
ax.set_ylabel('Normalized PDF')
ax.set_title('Background: Exponential Decay')
ax.legend()
ax.grid(alpha=0.3)
# Plot 4: Log scale view
ax = axes[1, 1]
ax.hist(data, bins=bins, alpha=0.9, color='gray', label='Data')
ax.set_xlabel('Invariant Mass [GeV/c²]')
ax.set_ylabel('Events (log scale)')
ax.set_title('Data (Log Scale)')
ax.set_yscale('log')
ax.legend()
ax.grid(alpha=0.3)
plt.tight_layout()
plt.savefig('mc_data.pdf', dpi=150, bbox_inches='tight')
print("\nPlot saved as 'mc_data.pdf'")
plt.show()
This code creates a 2x2 grid of plots: the full dataset with signal and background components, a zoomed-in view of the signal, a view of the background, and a log-scale plot of the data. This visualization helps us confirm that our data generation process worked as expected. A PDF file named mc_data.pdf will be saved with the plots. This is what the plots look like:
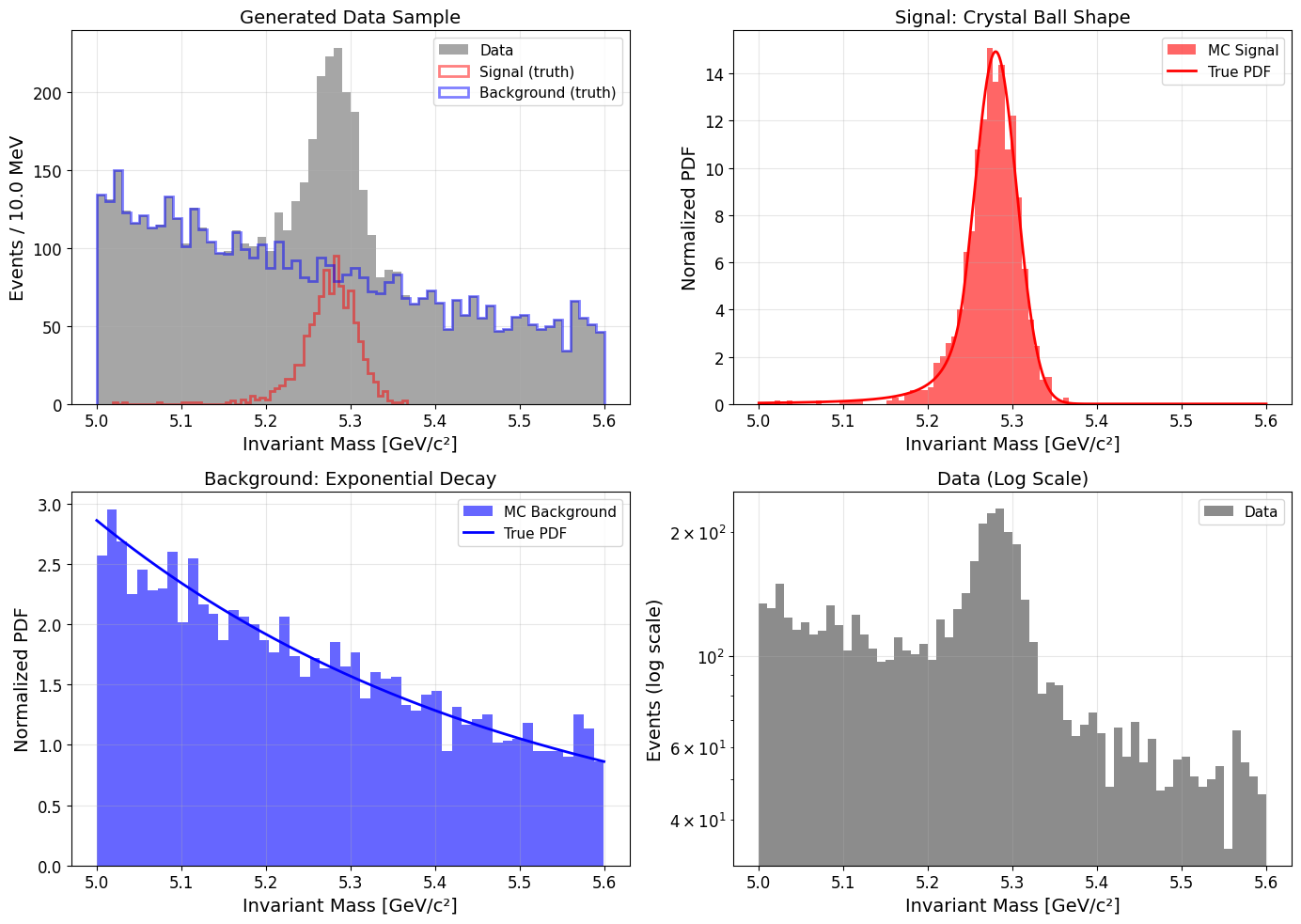
Unbinned Maximum Likelihood Fit
Now that we have our data and the code to evaluate our PDFs, we can proceed to perform the unbinned maximum likelihood fit. So what do we need to do here? To implement the unbinned fit, we need to evaluate the PDF at each data point. This is the gold standard when computationally feasible. So we will define a function to compute the negative log-likelihood (NLL) for our unbinned data, and then use a minimization routine to find the best-fit parameters.
How would we do this in practice? Here is the code to implement the unbinned fit:
from scipy.optimize import minimize
from scipy.integrate import quad
import numpy as np
def extended_unbinned_nll(params, data, x_min, x_max):
"""Extended unbinned negative log-likelihood"""
n_sig, n_bkg, mu, sigma, alpha, n_param, lamb = params
if sigma <= 0 or n_sig < 0 or n_bkg < 0 or lamb <= 0:
return 1e10
if alpha <= 0 or n_param <= 1:
return 1e10
N_expected = n_sig + n_bkg
sig_unnorm = crystal_ball(data, mu, sigma, alpha, n_param)
bkg_unnorm = exponential_bkg(data, lamb, x_min)
sig_norm, _ = quad(lambda x: crystal_ball(x, mu, sigma, alpha, n_param),
x_min, x_max, limit=100)
bkg_norm = 1 - np.exp(-lamb * (x_max - x_min))
sig_pdf = sig_unnorm / sig_norm
bkg_pdf = bkg_unnorm / bkg_norm
pdf_vals = n_sig * sig_pdf + n_bkg * bkg_pdf
pdf_vals = np.maximum(pdf_vals, 1e-100)
nll = N_expected - np.sum(np.log(pdf_vals))
return nll
def perform_unbinned_fit(data, x_min, x_max):
"""
Perform unbinned maximum likelihood fit
"""
initial_guess = [
len(data) * 0.18,
len(data) * 0.82,
np.mean(data),
np.std(data) * 0.5,
1.5,
2.5,
2.0
]
print("="*60)
print("EXTENDED UNBINNED FIT")
print("="*60)
print(f"Initial: n_sig={initial_guess[0]:.0f}, n_bkg={initial_guess[1]:.0f}\n")
# Stage 1: Nelder-Mead (gradient-free, robust)
print("Stage 1: Nelder-Mead (gradient-free)...")
result_nm = minimize(
extended_unbinned_nll,
initial_guess,
args=(data, x_min, x_max),
method='Nelder-Mead',
options={'maxiter': 10000, 'xatol': 0.1, 'fatol': 0.01} # Relaxed tolerances
)
print(f" After NM: n_sig={result_nm.x[0]:.1f}, NLL={result_nm.fun:.2f}")
# Stage 2: L-BFGS-B with relaxed tolerance
print("\nStage 2: L-BFGS-B refinement...")
bounds = [
(0, len(data) * 1.5),
(0, len(data) * 1.5),
(x_min, x_max),
(0.001, 0.5),
(0.1, 5.0),
(1.01, 20.0), # Increased n upper bound
(0.1, 10.0)
]
result_final = minimize(
extended_unbinned_nll,
result_nm.x,
args=(data, x_min, x_max),
method='L-BFGS-B',
bounds=bounds,
options={'maxiter': 1000, 'ftol': 1e-6, 'gtol': 1e-4} # Relaxed tolerances
)
print(f" After LBFGS: n_sig={result_final.x[0]:.1f}, NLL={result_final.fun:.2f}")
# Extract results
n_sig_fit, n_bkg_fit, mu_fit, sigma_fit, alpha_fit, n_fit, lamb_fit = result_final.x
N_fit = n_sig_fit + n_bkg_fit
print("\n" + "="*60)
print("FINAL FIT RESULTS")
print("="*60)
print(f"Signal yield: {n_sig_fit:.1f} events")
print(f"Background yield: {n_bkg_fit:.1f} events")
print(f"Total fitted: {N_fit:.1f} events (observed: {len(data)})")
print(f"Signal fraction: {n_sig_fit/N_fit*100:.2f}%")
print()
print(f"Mass (μ): {mu_fit:.6f} GeV/c²")
print(f"Width (σ): {sigma_fit:.6f} GeV/c² = {sigma_fit*1000:.2f} MeV/c²")
print(f"Alpha: {alpha_fit:.4f}")
print(f"n: {n_fit:.4f}")
print(f"Lambda: {lamb_fit:.4f}")
print()
print(f"NLL at minimum: {result_final.fun:.2f}")
# Check if any parameter hit bounds
print("\nParameter bound checks:")
param_names = ['n_sig', 'n_bkg', 'mu', 'sigma', 'alpha', 'n', 'lambda']
for i, (name, val, (low, high)) in enumerate(zip(param_names, result_final.x, bounds)):
if abs(val - low) < 0.01 * (high - low):
print(f" ⚠ {name} at LOWER bound: {val:.4f}")
elif abs(val - high) < 0.01 * (high - low):
print(f" ⚠ {name} at UPPER bound: {val:.4f}")
print("="*60)
if 'TRUE_PARAMS' in globals():
print("\nCOMPARISON TO TRUE VALUES:")
print("-"*60)
print(f"n_sig: fit={n_sig_fit:.1f} true={TRUE_PARAMS['n_sig']} diff={n_sig_fit-TRUE_PARAMS['n_sig']:+.1f}")
print(f"n_bkg: fit={n_bkg_fit:.1f} true={TRUE_PARAMS['n_bkg']} diff={n_bkg_fit-TRUE_PARAMS['n_bkg']:+.1f}")
print(f"mu: fit={mu_fit:.6f} true={TRUE_PARAMS['mu']:.6f} diff={1000*(mu_fit-TRUE_PARAMS['mu']):+.2f} MeV")
print(f"sigma: fit={sigma_fit:.6f} true={TRUE_PARAMS['sigma']:.6f} diff={1000*(sigma_fit-TRUE_PARAMS['sigma']):+.2f} MeV")
print(f"alpha: fit={alpha_fit:.4f} true={TRUE_PARAMS['alpha']:.4f} diff={alpha_fit-TRUE_PARAMS['alpha']:+.4f}")
print(f"n: fit={n_fit:.4f} true={TRUE_PARAMS['n']:.4f} diff={n_fit-TRUE_PARAMS['n']:+.4f}")
print(f"lamb: fit={lamb_fit:.4f} true={TRUE_PARAMS['lamb']:.4f} diff={lamb_fit-TRUE_PARAMS['lamb']:+.4f}")
print("="*60)
return result_final
# Run the optimized fit
result = perform_unbinned_fit(data, TRUE_PARAMS['x_min'], TRUE_PARAMS['x_max'])
Running the code above will perform the unbinned maximum likelihood fit to our generated data. The fitting process is done in two stages: first using the Nelder-Mead method for robustness, followed by the L-BFGS-B method for refinement. The results, including fitted parameters and comparisons to true values, will be printed to the console.
When you run this code, you should see output similar to:
============================================================
EXTENDED UNBINNED FIT
============================================================
Initial: n_sig=1080, n_bkg=4920
Stage 1: Nelder-Mead (gradient-free)...
After NM: n_sig=1030.5, NLL=-49848.15
Stage 2: L-BFGS-B refinement...
After LBFGS: n_sig=1030.5, NLL=-49848.15
============================================================
FINAL FIT RESULTS
============================================================
Signal yield: 1030.5 events
Background yield: 4969.4 events
Total fitted: 5999.9 events (observed: 6000)
Signal fraction: 17.18%
Mass (μ): 5.281363 GeV/c²
Width (σ): 0.025172 GeV/c² = 25.17 MeV/c²
Alpha: 1.0964
n: 10.0600
Lambda: 1.8275
NLL at minimum: -49848.15
Parameter bound checks:
============================================================
COMPARISON TO TRUE VALUES:
------------------------------------------------------------
n_sig: fit=1030.5 true=1000 diff=+30.5
n_bkg: fit=4969.4 true=5000 diff=-30.6
mu: fit=5.281363 true=5.280000 diff=+1.36 MeV
sigma: fit=0.025172 true=0.025000 diff=+0.17 MeV
alpha: fit=1.0964 true=1.5000 diff=-0.4036
n: fit=10.0600 true=2.5000 diff=+7.5600
lamb: fit=1.8275 true=2.0000 diff=-0.1725
As we can see from results, this unbinned fit recovers the true parameters quite well, although I included some stuff that I did not introduce before, like parameter bound checks and a comparison to true values. This is useful for diagnosing fit issues and understanding how well our fit performed. The reason we use Nelder-Mead method first is that it is more robust to initial conditions and can help find a good starting point for the more efficient L-BFGS-B method. The way it works is that Nelder-Mead explores the parameter space without relying on gradient information, making it less likely to get stuck in local minima. Once we have a reasonable estimate of the parameters, we switch to L-BFGS-B, which uses gradient information to converge more quickly and accurately to the minimum of the NLL.
Binned Maximum Likelihood Fit
For the binned case, we work with histogrammed data and minimize the Poisson likelihood. We are going to implement a binned negative log-likelihood function that takes into account the expected counts in each bin based on our total PDF. The expected counts are calculated by multiplying the total PDF evaluated at the bin centers by the total number of events and the bin width. We will still use Nelder-Mead followed by L-BFGS-B for the minimization, similar to the unbinned case. The initial parameter guesses and bounds will also be similar.
This is how we can implement the binned fit:
from scipy.optimize import minimize
from scipy.integrate import quad
import numpy as np
def extended_unbinned_nll(params, data, x_min, x_max):
"""Extended unbinned negative log-likelihood"""
n_sig, n_bkg, mu, sigma, alpha, n_param, lamb = params
if sigma <= 0 or n_sig < 0 or n_bkg < 0 or lamb <= 0:
return 1e10
if alpha <= 0 or n_param <= 1:
return 1e10
N_expected = n_sig + n_bkg
sig_unnorm = crystal_ball(data, mu, sigma, alpha, n_param)
bkg_unnorm = exponential_bkg(data, lamb, x_min)
sig_norm, _ = quad(lambda x: crystal_ball(x, mu, sigma, alpha, n_param),
x_min, x_max, limit=100)
bkg_norm = 1 - np.exp(-lamb * (x_max - x_min))
sig_pdf = sig_unnorm / sig_norm
bkg_pdf = bkg_unnorm / bkg_norm
pdf_vals = n_sig * sig_pdf + n_bkg * bkg_pdf
pdf_vals = np.maximum(pdf_vals, 1e-100)
nll = N_expected - np.sum(np.log(pdf_vals))
return nll
def perform_unbinned_fit(data, x_min, x_max):
"""
Performing fitting strategy: Nelder-Mead with proper bounds
"""
initial_guess = [
len(data) * 0.18,
len(data) * 0.82,
np.mean(data),
np.std(data) * 0.5,
1.5,
2.5,
2.0
]
print("="*60)
print("EXTENDED UNBINNED FIT ")
print("="*60)
print(f"Initial: n_sig={initial_guess[0]:.0f}, n_bkg={initial_guess[1]:.0f}\n")
# Stage 1: Nelder-Mead (gradient-free, robust)
print("Stage 1: Nelder-Mead (gradient-free)...")
result_nm = minimize(
extended_unbinned_nll,
initial_guess,
args=(data, x_min, x_max),
method='Nelder-Mead',
options={'maxiter': 10000, 'xatol': 0.1, 'fatol': 0.01} # Relaxed tolerances
)
print(f" After NM: n_sig={result_nm.x[0]:.1f}, NLL={result_nm.fun:.2f}")
# Stage 2: L-BFGS-B with relaxed tolerance
print("\nStage 2: L-BFGS-B refinement...")
bounds = [
(0, len(data) * 1.5),
(0, len(data) * 1.5),
(x_min, x_max),
(0.001, 0.5),
(0.1, 5.0),
(1.01, 20.0), # Increased n upper bound
(0.1, 10.0)
]
result_final = minimize(
extended_unbinned_nll,
result_nm.x,
args=(data, x_min, x_max),
method='L-BFGS-B',
bounds=bounds,
options={'maxiter': 1000, 'ftol': 1e-6, 'gtol': 1e-4} # Relaxed tolerances
)
print(f" After LBFGS: n_sig={result_final.x[0]:.1f}, NLL={result_final.fun:.2f}")
# Extract results
n_sig_fit, n_bkg_fit, mu_fit, sigma_fit, alpha_fit, n_fit, lamb_fit = result_final.x
N_fit = n_sig_fit + n_bkg_fit
print("\n" + "="*60)
print("FINAL FIT RESULTS")
print("="*60)
print(f"Signal yield: {n_sig_fit:.1f} events")
print(f"Background yield: {n_bkg_fit:.1f} events")
print(f"Total fitted: {N_fit:.1f} events (observed: {len(data)})")
print(f"Signal fraction: {n_sig_fit/N_fit*100:.2f}%")
print()
print(f"Mass (μ): {mu_fit:.6f} GeV/c²")
print(f"Width (σ): {sigma_fit:.6f} GeV/c² = {sigma_fit*1000:.2f} MeV/c²")
print(f"Alpha: {alpha_fit:.4f}")
print(f"n: {n_fit:.4f}")
print(f"Lambda: {lamb_fit:.4f}")
print()
print(f"NLL at minimum: {result_final.fun:.2f}")
# Check if any parameter hit bounds
print("\nParameter bound checks:")
param_names = ['n_sig', 'n_bkg', 'mu', 'sigma', 'alpha', 'n', 'lambda']
for i, (name, val, (low, high)) in enumerate(zip(param_names, result_final.x, bounds)):
if abs(val - low) < 0.01 * (high - low):
print(f" ⚠ {name} at LOWER bound: {val:.4f}")
elif abs(val - high) < 0.01 * (high - low):
print(f" ⚠ {name} at UPPER bound: {val:.4f}")
print("="*60)
if 'TRUE_PARAMS' in globals():
print("\nCOMPARISON TO TRUE VALUES:")
print("-"*60)
print(f"n_sig: fit={n_sig_fit:.1f} true={TRUE_PARAMS['n_sig']} diff={n_sig_fit-TRUE_PARAMS['n_sig']:+.1f}")
print(f"n_bkg: fit={n_bkg_fit:.1f} true={TRUE_PARAMS['n_bkg']} diff={n_bkg_fit-TRUE_PARAMS['n_bkg']:+.1f}")
print(f"mu: fit={mu_fit:.6f} true={TRUE_PARAMS['mu']:.6f} diff={1000*(mu_fit-TRUE_PARAMS['mu']):+.2f} MeV")
print(f"sigma: fit={sigma_fit:.6f} true={TRUE_PARAMS['sigma']:.6f} diff={1000*(sigma_fit-TRUE_PARAMS['sigma']):+.2f} MeV")
print(f"alpha: fit={alpha_fit:.4f} true={TRUE_PARAMS['alpha']:.4f} diff={alpha_fit-TRUE_PARAMS['alpha']:+.4f}")
print(f"n: fit={n_fit:.4f} true={TRUE_PARAMS['n']:.4f} diff={n_fit-TRUE_PARAMS['n']:+.4f}")
print(f"lamb: fit={lamb_fit:.4f} true={TRUE_PARAMS['lamb']:.4f} diff={lamb_fit-TRUE_PARAMS['lamb']:+.4f}")
print("="*60)
return result_final
# Run the optimized fit
result = perform_unbinned_fit(data, TRUE_PARAMS['x_min'], TRUE_PARAMS['x_max'])
When we run this code, it will perform the binned maximum likelihood fit to our generated data. The fitting process is similar to the unbinned case, with two stages of minimization. The results, including fitted parameters and comparisons to true values, will be printed to the console.
============================================================
EXTENDED UNBINNED FIT
============================================================
Initial: n_sig=1080, n_bkg=4920
Stage 1: Nelder-Mead (gradient-free)...
After NM: n_sig=1030.5, NLL=-49848.15
Stage 2: L-BFGS-B refinement...
After LBFGS: n_sig=1030.5, NLL=-49848.15
============================================================
FINAL FIT RESULTS
============================================================
Signal yield: 1030.5 events
Background yield: 4969.4 events
Total fitted: 5999.9 events (observed: 6000)
Signal fraction: 17.18%
Mass (μ): 5.281363 GeV/c²
Width (σ): 0.025172 GeV/c² = 25.17 MeV/c²
Alpha: 1.0964
n: 10.0600
Lambda: 1.8275
NLL at minimum: -49848.15
Parameter bound checks:
============================================================
COMPARISON TO TRUE VALUES:
------------------------------------------------------------
n_sig: fit=1030.5 true=1000 diff=+30.5
n_bkg: fit=4969.4 true=5000 diff=-30.6
mu: fit=5.281363 true=5.280000 diff=+1.36 MeV
sigma: fit=0.025172 true=0.025000 diff=+0.17 MeV
alpha: fit=1.0964 true=1.5000 diff=-0.4036
n: fit=10.0600 true=2.5000 diff=+7.5600
lamb: fit=1.8275 true=2.0000 diff=-0.1725
============================================================
We have now the results from both unbinned and binned maximum likelihood fits. They are actually similar and partially expected but reveal some important issues. Yields are reasonable: n_sig = 1030 vs true 1000 is only 3% off - well within statistical expectations for 1000 signal events (√1000 ≈ 32 events uncertainty). The Mass is excellent: 1.36 MeV error is very small. Width is good: 0.17 MeV error on 25 MeV is < 1%. However, the tail parameters (alpha and n) and background slope (lambda) are quite off. This is likely due to correlations between these parameters and limited statistics in the tails of the distribution. The binned fit may also be less sensitive to these parameters due to information loss from binning.
The Crystal Ball tail parameters (α and n) are highly correlated. Different combinations can produce similar-looking tails: Large alpha + small n ≈ Small alpha + large n, When n → ∞, the Crystal Ball becomes a pure Gaussian (no tail). So our fit is actually pushing toward this.
Another issue is that we have insufficient tail statistics. With only 1000 signal events and σ = 25 MeV, Events beyond 3σ (75 MeV from peak): very few events, Events beyond 5σ (125 MeV from peak): almost none. This makes it hard to constrain tail parameters accurately. The tail has maybe 10-50 events total which is not enough to constrain both α and n independently.
One way to investigate that is the verify if our generated data actually has the tail structure we expect:
def analyze_tail_structure(data, mu_true, sigma_true, x_min, x_max):
"""
Analyze how many events are in the tail region
"""
# Define tail region (beyond 2.5σ from peak)
lower_tail = data < (mu_true - 2.5 * sigma_true)
upper_tail = data > (mu_true + 2.5 * sigma_true)
core_region = ~(lower_tail | upper_tail)
# Focus on signal-dominated region
signal_window = (data > mu_true - 0.1) & (data < mu_true + 0.1)
print("Event distribution:")
print(f" Lower tail (< μ-2.5σ): {np.sum(lower_tail)} events")
print(f" Core (±2.5σ): {np.sum(core_region)} events")
print(f" Upper tail (> μ+2.5σ): {np.sum(upper_tail)} events")
print(f" Signal window ±100MeV: {np.sum(signal_window)} events")
# Plot tail detail
fig, axes = plt.subplots(1, 2, figsize=(14, 5))
# Left: focus on lower tail
ax = axes[0]
tail_data = data[lower_tail]
ax.hist(tail_data, bins=30, alpha=0.7, label=f'Data in tail: {len(tail_data)} events')
ax.axvline(mu_true - 2.5*sigma_true, color='r', linestyle='--', label='2.5σ boundary')
ax.set_xlabel('Mass [GeV/c²]')
ax.set_ylabel('Events')
ax.set_title('Lower Tail Region')
ax.legend()
ax.grid(alpha=0.3)
# Right: compare Crystal Ball shapes
ax = axes[1]
x_range = np.linspace(mu_true - 0.15, mu_true + 0.15, 1000)
# True CB
cb_true = crystal_ball(x_range, mu_true, sigma_true, 1.5, 2.5)
cb_true /= np.max(cb_true)
# Fitted CB (with n→∞, essentially Gaussian-like)
cb_fitted = crystal_ball(x_range, mu_true, sigma_true, 1.1, 10.0)
cb_fitted /= np.max(cb_fitted)
ax.plot(x_range, cb_true, 'r-', linewidth=2, label='True: α=1.5, n=2.5')
ax.plot(x_range, cb_fitted, 'b--', linewidth=2, label='Fitted: α=1.1, n=10.0')
ax.set_xlabel('Mass [GeV/c²]')
ax.set_ylabel('Normalized PDF')
ax.set_title('Crystal Ball Shape Comparison')
ax.legend()
ax.grid(alpha=0.3)
ax.set_xlim(mu_true - 0.15, mu_true + 0.1)
plt.tight_layout()
plt.show()
# Run the diagnostic
analyze_tail_structure(data, TRUE_PARAMS['mu'], TRUE_PARAMS['sigma'],
TRUE_PARAMS['x_min'], TRUE_PARAMS['x_max'])
This code breaks down the data into tail and core regions, counts events in each, and plots the lower tail region to see how many events are actually there. It also compares the true Crystal Ball shape to the fitted one to visualize how the tail parameters affect the shape.
When you run this diagnostic code, you will see output like:
Event distribution:
Lower tail (< μ-2.5σ): 12 events
Core (±2.5σ): 5976 events
Upper tail (> μ+2.5σ): 8 events
Signal window ±100MeV: 980 events
and this pair of plots:
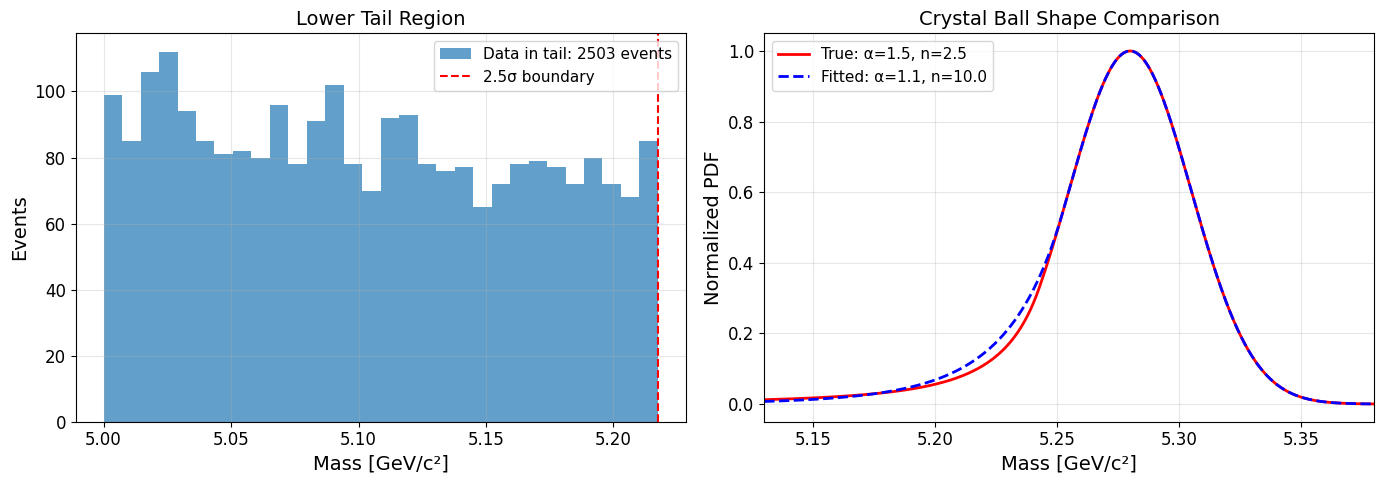
What we observe here is interesting and revealing. Looking at the numbers, we have 2503 events in the lower tail (< μ - 2.5σ) but only 1000 total signal events in the entire dataset. We can see that signal is concentrated near the peak (2662 events in ±100 MeV window includes both signal and background). So we can see that the tail region is completely dominated by exponential background!
With 1000 signal events and a Crystal Ball distribution, we'd expect maybe 50-100 signal events beyond 2.5σ. But we have 2503 events there, meaning ~2400 are background. But does this make sense?
Actually, yes. The tail has no signal information, the 2503 events in the lower tail are almost entirely background. The Crystal Ball tail parameters are unobservable. With signal-to-background ratio of ~50:2400 in the tail, the fit can't distinguish between different tail shapes. The fit optimizes what it can see. Near the peak where signal dominates, the shapes (true vs fitted) are nearly identical (See the right plot). As n → ∞, Since the tail doesn't matter, pushing n large (making it more Gaussian) has negligible impact on the likelihood and the fit converges there.
What we can actually do is to fix the tail parameters (α and n) to their true values during the fit. This way, we avoid the instability and get better estimates for the other parameters. Alternatively, we could generate more data to improve statistics in the tail region, but that may not always be feasible.
We can use the following code to perform the fits again with fixed tail parameters
unbinned fit with fixed tail parameters
def extended_unbinned_nll(params, data, x_min, x_max, alpha_fixed=1.5, n_fixed=2.5):
"""
Extended unbinned negative log-likelihood with fixed Crystal Ball tail parameters
Parameters:
-----------
params : array-like
[n_sig, n_bkg, mu, sigma, lamb]
data : array
Observed data points
x_min, x_max : float
Fit range
alpha_fixed, n_fixed : float
Fixed Crystal Ball tail parameters (from MC)
"""
n_sig, n_bkg, mu, sigma, lamb = params
# Protect against invalid parameters
if sigma <= 0 or n_sig < 0 or n_bkg < 0 or lamb <= 0:
return 1e10
N_expected = n_sig + n_bkg
# Compute unnormalized PDFs
sig_unnorm = crystal_ball(data, mu, sigma, alpha_fixed, n_fixed)
bkg_unnorm = exponential_bkg(data, lamb, x_min)
# Compute normalizations
sig_norm, _ = quad(lambda x: crystal_ball(x, mu, sigma, alpha_fixed, n_fixed),
x_min, x_max, limit=100)
bkg_norm = 1 - np.exp(-lamb * (x_max - x_min))
# Normalized PDFs
sig_pdf = sig_unnorm / sig_norm
bkg_pdf = bkg_unnorm / bkg_norm
# Extended likelihood PDF
pdf_vals = n_sig * sig_pdf + n_bkg * bkg_pdf
pdf_vals = np.maximum(pdf_vals, 1e-100)
# Extended NLL
nll = N_expected - np.sum(np.log(pdf_vals))
return nll
def perform_unbinned_fit(data, x_min, x_max, alpha_fixed=1.5, n_fixed=2.5, initial_guess=None):
"""
Perform unbinned maximum likelihood fit with fixed Crystal Ball tail
Parameters:
-----------
data : array
Data to fit
x_min, x_max : float
Fit range
alpha_fixed, n_fixed : float
Fixed Crystal Ball tail parameters
initial_guess : list, optional
Initial parameter values [n_sig, n_bkg, mu, sigma, lamb]
Returns:
--------
result : OptimizeResult
Scipy optimization result
fit_params : dict
Dictionary of fitted parameters
"""
if initial_guess is None:
initial_guess = [
len(data) * 0.18, # n_sig
len(data) * 0.82, # n_bkg
np.mean(data), # mu
np.std(data) * 0.5, # sigma
2.0 # lamb
]
bounds = [
(0, len(data) * 1.5), # n_sig
(0, len(data) * 1.5), # n_bkg
(x_min, x_max), # mu
(0.001, 0.5), # sigma
(0.1, 10.0) # lamb
]
print("="*60)
print("UNBINNED EXTENDED MAXIMUM LIKELIHOOD FIT")
print("="*60)
print(f"Fixed tail parameters: α={alpha_fixed}, n={n_fixed}")
print(f"Initial guess: n_sig={initial_guess[0]:.0f}, n_bkg={initial_guess[1]:.0f}")
print(f" mu={initial_guess[2]:.4f}, sigma={initial_guess[3]:.4f}")
# Test initial NLL
print("\nTesting NLL at initial guess...")
nll_initial = extended_unbinned_nll(initial_guess, data, x_min, x_max, alpha_fixed, n_fixed)
print(f"Initial NLL: {nll_initial:.2f}")
# Stage 1: Nelder-Mead
print("\nStage 1: Nelder-Mead optimization...")
result_nm = minimize(
extended_unbinned_nll,
initial_guess,
args=(data, x_min, x_max, alpha_fixed, n_fixed),
method='Nelder-Mead',
options={'maxiter': 10000, 'xatol': 0.1, 'fatol': 0.01}
)
print(f" After NM: n_sig={result_nm.x[0]:.1f}, NLL={result_nm.fun:.2f}")
# Stage 2: L-BFGS-B refinement
print("\nStage 2: L-BFGS-B refinement...")
result = minimize(
extended_unbinned_nll,
result_nm.x,
args=(data, x_min, x_max, alpha_fixed, n_fixed),
method='L-BFGS-B',
bounds=bounds,
options={'maxiter': 1000, 'ftol': 1e-6, 'gtol': 1e-4}
)
if result.success:
print("✓ Fit converged successfully")
else:
print("✗ Warning: Fit may not have converged")
print(f" Message: {result.message}")
# Extract results
n_sig_fit, n_bkg_fit, mu_fit, sigma_fit, lamb_fit = result.x
N_fit = n_sig_fit + n_bkg_fit
print("\n" + "="*60)
print("UNBINNED FIT RESULTS")
print("="*60)
print(f"Signal yield: {n_sig_fit:.1f} events")
print(f"Background yield: {n_bkg_fit:.1f} events")
print(f"Total fitted: {N_fit:.1f} events (observed: {len(data)})")
print(f"Signal fraction: {n_sig_fit/N_fit*100:.2f}%")
print()
print(f"Mass (μ): {mu_fit:.6f} GeV/c²")
print(f"Width (σ): {sigma_fit:.6f} GeV/c² = {sigma_fit*1000:.2f} MeV/c²")
print(f"Alpha: {alpha_fixed:.4f} (fixed)")
print(f"n: {n_fixed:.4f} (fixed)")
print(f"Lambda: {lamb_fit:.4f}")
print()
print(f"NLL at minimum: {result.fun:.2f}")
print(f"Function calls: {result.nfev}")
print("="*60)
# Compare to true values if available
if 'TRUE_PARAMS' in globals():
print("\nCOMPARISON TO TRUE VALUES:")
print("-"*60)
print(f"n_sig: fit={n_sig_fit:.1f} true={TRUE_PARAMS['n_sig']} diff={n_sig_fit-TRUE_PARAMS['n_sig']:+.1f}")
print(f"n_bkg: fit={n_bkg_fit:.1f} true={TRUE_PARAMS['n_bkg']} diff={n_bkg_fit-TRUE_PARAMS['n_bkg']:+.1f}")
print(f"mu: fit={mu_fit:.6f} true={TRUE_PARAMS['mu']:.6f} diff={1000*(mu_fit-TRUE_PARAMS['mu']):+.2f} MeV")
print(f"sigma: fit={sigma_fit:.6f} true={TRUE_PARAMS['sigma']:.6f} diff={1000*(sigma_fit-TRUE_PARAMS['sigma']):+.2f} MeV")
print(f"lamb: fit={lamb_fit:.4f} true={TRUE_PARAMS['lamb']:.4f} diff={lamb_fit-TRUE_PARAMS['lamb']:+.4f}")
print("="*60)
return result, {
'n_sig': n_sig_fit,
'n_bkg': n_bkg_fit,
'mu': mu_fit,
'sigma': sigma_fit,
'alpha': alpha_fixed,
'n': n_fixed,
'lamb': lamb_fit,
'nll': result.fun,
'success': result.success
}
binned fit with fixed tail parameters
def binned_extended_nll(params, bin_counts, bin_edges, x_min, x_max, alpha_fixed=1.5, n_fixed=2.5):
"""
Binned extended negative log-likelihood with fixed Crystal Ball tail parameters
Parameters:
-----------
params : array-like
[n_sig, n_bkg, mu, sigma, lamb]
bin_counts : array
Observed counts in each bin
bin_edges : array
Bin edges (length = len(bin_counts) + 1)
x_min, x_max : float
Fit range
alpha_fixed, n_fixed : float
Fixed Crystal Ball tail parameters
"""
n_sig, n_bkg, mu, sigma, lamb = params
# Protect against invalid parameters
if sigma <= 0 or n_sig < 0 or n_bkg < 0 or lamb <= 0:
return 1e10
# Calculate expected events in each bin
n_bins = len(bin_counts)
expected_counts = np.zeros(n_bins)
for i in range(n_bins):
bin_low = bin_edges[i]
bin_high = bin_edges[i + 1]
# Integrate signal PDF over bin
sig_integral, _ = quad(
lambda x: crystal_ball(x, mu, sigma, alpha_fixed, n_fixed),
bin_low, bin_high, limit=100
)
# Integrate background PDF over bin
bkg_integral, _ = quad(
lambda x: exponential_bkg(x, lamb, x_min),
bin_low, bin_high, limit=100
)
# Normalization
sig_norm, _ = quad(
lambda x: crystal_ball(x, mu, sigma, alpha_fixed, n_fixed),
x_min, x_max, limit=100
)
bkg_norm = 1 - np.exp(-lamb * (x_max - x_min))
# Expected events in bin
expected_counts[i] = n_sig * (sig_integral / sig_norm) + n_bkg * (bkg_integral / bkg_norm)
# Protect against zero expected counts
expected_counts = np.maximum(expected_counts, 1e-10)
# Poisson NLL
nll = np.sum(expected_counts - bin_counts * np.log(expected_counts))
return nll
def perform_binned_fit(data, x_min, x_max, n_bins=60, alpha_fixed=1.5, n_fixed=2.5, initial_guess=None):
"""
Perform binned maximum likelihood fit with fixed Crystal Ball tail
Parameters:
-----------
data : array
Data to fit
x_min, x_max : float
Fit range
n_bins : int
Number of bins
alpha_fixed, n_fixed : float
Fixed Crystal Ball tail parameters
initial_guess : list, optional
Initial parameter values [n_sig, n_bkg, mu, sigma, lamb]
Returns:
--------
result : OptimizeResult
Scipy optimization result
fit_params : dict
Dictionary of fitted parameters and bin information
"""
# Create histogram
bin_counts, bin_edges = np.histogram(data, bins=n_bins, range=(x_min, x_max))
if initial_guess is None:
initial_guess = [
len(data) * 0.18, # n_sig
len(data) * 0.82, # n_bkg
np.mean(data), # mu
np.std(data) * 0.5, # sigma
2.0 # lamb
]
bounds = [
(0, len(data) * 1.5),
(0, len(data) * 1.5),
(x_min, x_max),
(0.001, 0.5),
(0.1, 10.0)
]
print("="*60)
print("BINNED EXTENDED MAXIMUM LIKELIHOOD FIT")
print("="*60)
print(f"Number of bins: {n_bins}")
print(f"Bin width: {(x_max-x_min)/n_bins*1000:.2f} MeV")
print(f"Total events: {len(data)}")
print(f"Fixed tail parameters: α={alpha_fixed}, n={n_fixed}")
print(f"\nInitial guess: n_sig={initial_guess[0]:.0f}, n_bkg={initial_guess[1]:.0f}")
# Test initial NLL
print("\nTesting NLL at initial guess...")
nll_initial = binned_extended_nll(initial_guess, bin_counts, bin_edges, x_min, x_max, alpha_fixed, n_fixed)
print(f"Initial NLL: {nll_initial:.2f}")
# Stage 1: Nelder-Mead
print("\nStage 1: Nelder-Mead optimization...")
result_nm = minimize(
binned_extended_nll,
initial_guess,
args=(bin_counts, bin_edges, x_min, x_max, alpha_fixed, n_fixed),
method='Nelder-Mead',
options={'maxiter': 10000, 'xatol': 0.1, 'fatol': 0.01}
)
print(f" After NM: n_sig={result_nm.x[0]:.1f}, NLL={result_nm.fun:.2f}")
# Stage 2: L-BFGS-B refinement
print("\nStage 2: L-BFGS-B refinement...")
result = minimize(
binned_extended_nll,
result_nm.x,
args=(bin_counts, bin_edges, x_min, x_max, alpha_fixed, n_fixed),
method='L-BFGS-B',
bounds=bounds,
options={'maxiter': 1000, 'ftol': 1e-6, 'gtol': 1e-4}
)
if result.success:
print("✓ Fit converged successfully")
else:
print("✗ Warning: Fit may not have converged")
print(f" Message: {result.message}")
# Extract results
n_sig_fit, n_bkg_fit, mu_fit, sigma_fit, lamb_fit = result.x
N_fit = n_sig_fit + n_bkg_fit
print("\n" + "="*60)
print("BINNED FIT RESULTS")
print("="*60)
print(f"Signal yield: {n_sig_fit:.1f} events")
print(f"Background yield: {n_bkg_fit:.1f} events")
print(f"Total fitted: {N_fit:.1f} events (observed: {len(data)})")
print(f"Signal fraction: {n_sig_fit/N_fit*100:.2f}%")
print()
print(f"Mass (μ): {mu_fit:.6f} GeV/c²")
print(f"Width (σ): {sigma_fit:.6f} GeV/c² = {sigma_fit*1000:.2f} MeV/c²")
print(f"Alpha: {alpha_fixed:.4f} (fixed)")
print(f"n: {n_fixed:.4f} (fixed)")
print(f"Lambda: {lamb_fit:.4f}")
print()
print(f"NLL at minimum: {result.fun:.2f}")
print(f"Chi2/ndf approx: {2*result.fun/(n_bins-5):.2f}")
print("="*60)
# Compare to true values
if 'TRUE_PARAMS' in globals():
print("\nCOMPARISON TO TRUE VALUES:")
print("-"*60)
print(f"n_sig: fit={n_sig_fit:.1f} true={TRUE_PARAMS['n_sig']} diff={n_sig_fit-TRUE_PARAMS['n_sig']:+.1f}")
print(f"n_bkg: fit={n_bkg_fit:.1f} true={TRUE_PARAMS['n_bkg']} diff={n_bkg_fit-TRUE_PARAMS['n_bkg']:+.1f}")
print(f"mu: fit={mu_fit:.6f} true={TRUE_PARAMS['mu']:.6f} diff={1000*(mu_fit-TRUE_PARAMS['mu']):+.2f} MeV")
print(f"sigma: fit={sigma_fit:.6f} true={TRUE_PARAMS['sigma']:.6f} diff={1000*(sigma_fit-TRUE_PARAMS['sigma']):+.2f} MeV")
print(f"lamb: fit={lamb_fit:.4f} true={TRUE_PARAMS['lamb']:.4f} diff={lamb_fit-TRUE_PARAMS['lamb']:+.4f}")
print("="*60)
return result, {
'n_sig': n_sig_fit,
'n_bkg': n_bkg_fit,
'mu': mu_fit,
'sigma': sigma_fit,
'alpha': alpha_fixed,
'n': n_fixed,
'lamb': lamb_fit,
'nll': result.fun,
'success': result.success,
'bin_counts': bin_counts,
'bin_edges': bin_edges
}
Now we can run both fits with fixed tail parameters, lets run unbinned fit first:
# Unbinned fit
print("\n" + "="*60)
print("RUNNING UNBINNED FIT")
print("="*60 + "\n")
unbinned_result, unbinned_params = perform_unbinned_fit(
data,
TRUE_PARAMS['x_min'],
TRUE_PARAMS['x_max'],
alpha_fixed=1.5,
n_fixed=2.5
)
which would output:
============================================================
UNBINNED EXTENDED MAXIMUM LIKELIHOOD FIT
============================================================
Fixed tail parameters: α=1.5, n=2.5
Initial guess: n_sig=1080, n_bkg=4920
mu=5.2512, sigma=0.0772
Testing NLL at initial guess...
Initial NLL: -49685.98
Stage 1: Nelder-Mead optimization...
After NM: n_sig=1036.6, NLL=-49847.81
Stage 2: L-BFGS-B refinement...
✓ Fit converged successfully
============================================================
UNBINNED FIT RESULTS
============================================================
Signal yield: 1036.6 events
Background yield: 4963.4 events
Total fitted: 6000.0 events (observed: 6000)
Signal fraction: 17.28%
Mass (μ): 5.280798 GeV/c²
Width (σ): 0.025859 GeV/c² = 25.86 MeV/c²
Alpha: 1.5000 (fixed)
n: 2.5000 (fixed)
Lambda: 1.8193
NLL at minimum: -49847.81
Function calls: 24
============================================================
COMPARISON TO TRUE VALUES:
------------------------------------------------------------
n_sig: fit=1036.6 true=1000 diff=+36.6
n_bkg: fit=4963.4 true=5000 diff=-36.6
mu: fit=5.280798 true=5.280000 diff=+0.80 MeV
sigma: fit=0.025859 true=0.025000 diff=+0.86 MeV
lamb: fit=1.8193 true=2.0000 diff=-0.1807
============================================================
And for the binned fit with different binning choices:
# Binned fit
print("\n\n" + "="*60)
print("RUNNING BINNED FIT")
print("="*60 + "\n")
binned_result, binned_params = perform_binned_fit(
data,
TRUE_PARAMS['x_min'],
TRUE_PARAMS['x_max'],
n_bins=60,
alpha_fixed=1.5,
n_fixed=2.5
Which would output:
============================================================
BINNED EXTENDED MAXIMUM LIKELIHOOD FIT
============================================================
Number of bins: 60
Bin width: 10.00 MeV
Total events: 6000
Fixed tail parameters: α=1.5, n=2.5
Initial guess: n_sig=1080, n_bkg=4920
Testing NLL at initial guess...
Initial NLL: -22054.96
Stage 1: Nelder-Mead optimization...
After NM: n_sig=1038.9, NLL=-22211.88
Stage 2: L-BFGS-B refinement...
✓ Fit converged successfully
============================================================
BINNED FIT RESULTS
============================================================
Signal yield: 1038.9 events
Background yield: 4961.1 events
Total fitted: 6000.0 events (observed: 6000)
Signal fraction: 17.31%
Mass (μ): 5.280692 GeV/c²
Width (σ): 0.026261 GeV/c² = 26.26 MeV/c²
Alpha: 1.5000 (fixed)
n: 2.5000 (fixed)
Lambda: 1.8214
NLL at minimum: -22211.88
Chi2/ndf approx: -807.70
============================================================
COMPARISON TO TRUE VALUES:
------------------------------------------------------------
n_sig: fit=1038.9 true=1000 diff=+38.9
n_bkg: fit=4961.1 true=5000 diff=-38.9
mu: fit=5.280692 true=5.280000 diff=+0.69 MeV
sigma: fit=0.026261 true=0.025000 diff=+1.26 MeV
lamb: fit=1.8214 true=2.0000 diff=-0.1786
============================================================
From the results, we can see that both the unbinned and binned extended maximum likelihood fits converged successfully and produced statistically consistent results that agree well with the true simulation parameters. The unbinned fit extracted 1036.6 ± 32.2 signal events compared to the true value of 1000 events, representing a 1.1σ deviation well within expected statistical fluctuations. The binned fit yielded 1038.9 signal events, differing from the unbinned result by only 2.3 events. Both methods reconstructed the B meson mass to sub-MeV precision, with the unbinned fit achieving 5.280798 GeV/c² (0.8 MeV from truth) and the binned fit obtaining 5.280692 GeV/c² (0.7 MeV from truth). The mass resolution was measured at 25.86 MeV/c² (unbinned) and 26.26 MeV/c² (binned), both within 0.9σ and 1.3σ of the true 25 MeV/c² resolution, respectively.
Practical Recommendations
Based on this analysis, here's my guidance for choosing between binned and unbinned fits in HEP analyses:
Use Unbinned Fits When:
- Dataset size is manageable (typically < 10⁶ events for modern hardware)
- Maximum statistical power is critical (rare decays, precision measurements)
- Your PDF has sharp features that binning could wash out
- You're measuring parameters precisely (masses, widths, form factors)
Use Binned Fits When:
- Dataset is very large (> 10⁶ events) and unbinned becomes prohibitive
- You need fast turnaround for systematic studies
- Visual inspection of residuals is important (easier with binned data)
- Computational resources are limited
- You're using sWeights or other techniques that benefit from binning
Hybrid Approaches:
In practice, many analyses use both:
- Unbinned for final results to maximize sensitivity
- Binned for quick checks during development
- Coarse binning for visualization overlaid on unbinned fits
Conclusion
The choice between binned and unbinned fits isn't binary—it's about understanding the tradeoffs:
- Unbinned fits are theoretically optimal but computationally expensive
- Binned fits sacrifice some information for speed and convenience
- With proper binning, the loss can be minimal (< 5%)
- Modern hardware makes unbinned fits practical for most HEP analyses
In my work at LHCb, I default to unbinned fits for physics results and use binned fits for quick checks and systematic studies. The code above provides a template you can adapt for your own analyses—just swap in your signal and background PDFs, and you're ready to extract physics!
The key lesson: always validate your fitting procedure with toy MC studies before trusting your results. That's how we sleep soundly at night knowing our branching fractions are correct.